From microfluidics to metastasis
New platform enables longitudinal studies of circulating tumor cells in mouse models of cancer.
Circulating tumor cells (CTCs) — an intermediate form of cancer cell between a primary and metastatic tumor cell — carry a treasure trove of information that is critical to treating cancer. Numerous engineering advancements over the years have made it possible to extract cells via liquid biopsy and analyze them to monitor an individual patient’s response to treatment and predict relapse.
Thanks to significant progress toward creating genetically engineered mouse models, liquid biopsies hold great promise for the lab as well. These mouse models mimic many aspects of human tumor development and have enabled informative studies that cannot be performed in patients. For example, these models can be used to trace the evolution of cells from initial mutation to eventual metastasis, a process in which CTCs play a critical role. But since it has not been possible to monitor CTCs over time in mice, scientists’ ability to study important features of metastasis has been limited.
The challenge lies in capturing enough cells to conduct such longitudinal studies. Although primary tumors shed CTCs constantly, the density of CTCs in blood is very low — fewer than 100 CTCs per milliliter. For human patients undergoing liquid biopsy, this does not present a problem. Clinicians can withdraw enough blood to guarantee a sufficient sample of CTCs, just a few milliliters out of five or so liters on average, with minimal impact to the patient.
A mouse, on the other hand, only has about 1.5 milliliters of blood in total. If researchers want to study CTCs over time, they may safely withdraw no more than a few microliters of blood from a mouse each day — nowhere near enough to ensure that many (or any) CTCs are collected.
But with a new approach developed by researchers at the Koch Institute for Integrative Cancer Research, it is now possible to collect CTCs from mice over days and even weeks, and analyze them as the disease progresses. The system, described in the Proceedings of the National Academy of Sciences the week of Jan. 21, diverts blood to a microfluidic cell-sorting chip that extracts individual CTCs before returning the blood back to an awake mouse.
A menu of sorts
The inspiration for the project was cooked up, not in the lab, but during a chance encounter in the Koch Café between Tyler Jacks, director of the Koch Institute and the David H. Koch Professor of Biology, and Scott Manalis, the Andrew and Erna Viterbi Professor in the departments of Biological Engineering and Mechanical Engineering and a member of the Koch Institute.
As luck and lunch lines would have it, the pair would discuss thesis work being done by then-graduate student Shawn Davidson, who was using a dialysis-like system to track metabolites in the bloodstream of mice in the laboratory of Matthew Vander Heiden, an associate professor of biology. Jacks and Manalis were inspired: Could a similar approach could be used to study rare CTCs in real time?
Along with their Koch Institute colleague Alex K. Shalek, the Pfizer-Laubach Career Development Assistant Professor of Chemistry and a core member of the Institute for Medical Engineering and Science (IMES) at MIT, it would take Jacks and Manalis more than five years to put all the pieces of the system together, drawing from different areas of expertise around the Koch Institute. The Jacks lab supplied its fluorescent small cell lung cancer model, the Manalis lab developed the real-time CTC isolation platform, and the Shalek lab provided genomic profiles of the collected CTCs using single-cell RNA sequencing.
“This is a project that could not have succeeded without a sustained effort from several labs with very different sets of expertise. For my lab, which primarily consists of engineers, the opportunity to participate in this type of research has been incredibly exciting and is the reason why we are in the Koch Institute,” Manalis says.
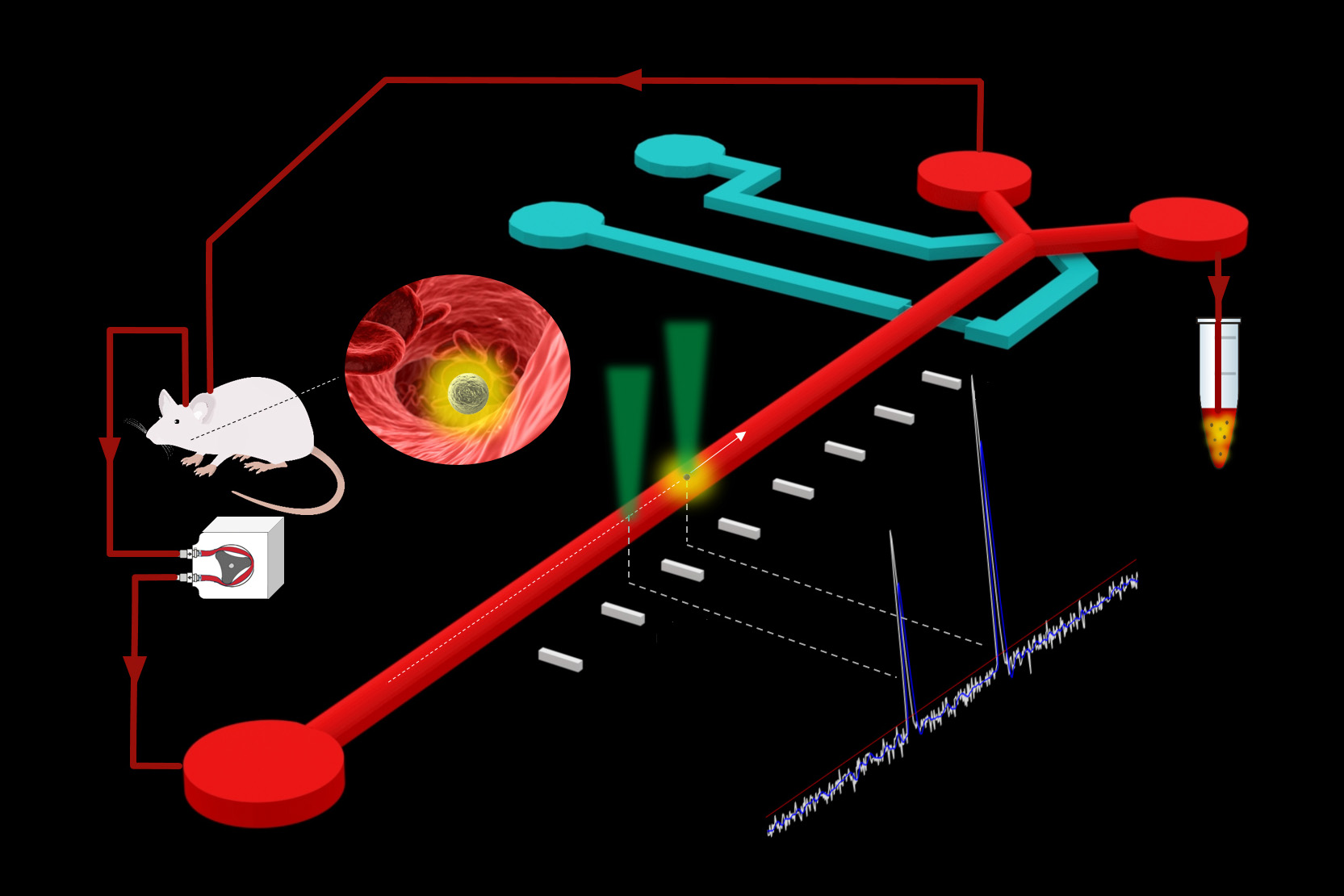
The CTC sorter uses laser excitation to identify tumor cells expressing a fluorescent marker that is incorporated in the mouse model. The system draws blood from the mouse and passes it through a microfluidic chip to detect and extract the fluorescing CTCs before returning the blood back to the mouse. A minute amount of blood — approximately 100 nanoliters — is diverted with every detected CTC into a collection tube, which then is purified further to extract individual CTCs from the thousands of other blood cells.
“The real-time detection of CTCs happens at a flow rate of approximately 2 milliliters per hour which allows us to scan nearly the entire blood volume of an awake and moving mouse within an hour,” says Bashar Hamza, a graduate student in the Manalis lab and one of the lead authors on the paper.
Biology in their blood
With the development of a real-time cell sorter, the researchers could now, for the first time, longitudinally collect CTCs from the same mouse.
Previously, the low blood volume of mice and the rarity of CTCs meant that groups of mice had to be sacrificed at successive times so that their CTCs could be pooled. However, CTCs from different mice often have significantly different gene expression profiles that can obscure subtle changes that occur from the evolution of the tumor or a perturbation such as a drug.
To demonstrate that their cell-sorter could capture these differences, the researchers treated mice with a compound called JQ1, which is known to inhibit the proliferation of cancer cells and perturb gene expression. CTCs were collected and profiled with single-cell RNA sequencing for two hours prior to the treatment, and then every 24 hours after the initial treatment for four days.
When the researchers pooled data for all mice that had been treated with JQ1, they found that the data clustered based on individual mice, offering no confirmation that the drug affects CTC gene expression over time. However, when the researchers analyzed single-mouse data, they observed gene expression shift with time.
“What’s so exciting about this platform and our approach is that we finally have the opportunity to comprehensively study longitudinal CTC responses without worrying about the potentially confounding effects of mouse-to-mouse variability. I, for one, can’t wait to see what we will be able to learn as we profile more CTCs, and their matched primary and metastatic tumors,” says Shalek.
Researchers believe their approach, which they intend to use in additional cancer types including non-small cell lung, pancreatic, and breast cancer, could open new avenues of inquiry in the study of CTCs, such as studying long-term drug responses, characterizing their relationship to metastatic tumors, and measuring their rate of production in short timeframes — and the entire metastatic cascade. In future work, researchers also plan to use their approach for profiling rare immune cells and monitoring cells in dynamic contexts such as wound healing and tumor formation.
“The ability to study CTCs as well other rare cells in the blood longitudinally gives us a powerful view into cancer development. This sorter represents a real breakthrough for the field and it is a great example of the Koch Institute in action,” says Jacks.
The paper’s other co-lead authors are graduate students Sheng Rong Ng from the Jacks lab and Sanjay Prakadan from the Shalek lab. The research is supported, in part, by the Ludwig Center at MIT, the National Cancer Institute, the National Institutes of Health and the Searle Scholars Program.