Building a platform to image membrane proteins
Biologists devise an efficient method to prepare fluorescently tagged proteins and simulate their native environment.
All cells have a lipid membrane that encircles their internal components — forming a protective barrier to control what gets in and what stays out. The proteins embedded in these membranes are essential for life; they help facilitate nutrient transport, energy conversion and storage, and cellular communication. They are also important in human disease, and represent around 60 percent of approved drug targets. In order to study these membrane proteins outside the complexity of the cell, researchers must use detergent to strip away the membrane and extract them. However, determining the best detergent for each protein can involve extensive trial and error. And, removing a protein from its natural environment risks destabilizing the folded structure and disrupting function.
In a study published on Dec. 9 in Cell Chemical Biology, scientists from MIT devised a rapid and generalizable way to extract, purify, and label membrane proteins for imaging without any detergent at all — bringing along a portion of the surrounding membrane to protect the protein and simulate its natural environment. Their approach combines well-established chemical and biochemical techniques in a new way, efficiently isolating the protein so it can be fluorescently labeled and examined under a microscope.
“I always joke that it’s not very lifelike to study proteins in soap,” says senior author Barbara Imperiali, a professor of biology and chemistry. “We’ve created a workflow that allows membrane proteins to be imaged while maintaining their native identities and interactions. Hopefully now fewer people will shy away from studying membrane proteins, given their importance in many physiological processes.”
As a member of the Imperiali lab, former postdoc and lead author Jean-Marie Swiecicki investigated membrane proteins from the foodborne pathogen Campylobacter jejuni. In this study, Swiecicki focused on PglC and PglA, two membrane proteins that play a role in enabling the bacteria to infect human cells. His experiments required labeling PglC and PglA with fluorescent tags in order to track them. However, he wasn’t satisfied with existing methods to do so.
In some cases, the fluorescent tags that must be incorporated into the protein in order to visualize it are too large to be placed at defined positions. In other cases, these tags don’t shine brightly enough, or interfere with the structure and function of the protein.
To avoid such issues, Swiecicki decided to use a method known as “unnatural amino-acid mutagenesis.” Amino acids are the units that compose the protein, and unnatural amino-acid mutagenesis involves adding a new amino acid containing an engineered chemical group within the protein sequence. This chemical group can then be labeled with a brightly glowing tag.
Swiecicki inserted the genetic code for the C. jejuni membrane proteins into a different bacterium, Escherichia coli. Inside E. coli, he could incorporate the unnatural amino acid, which could be chemically modified to add the fluorescent label.
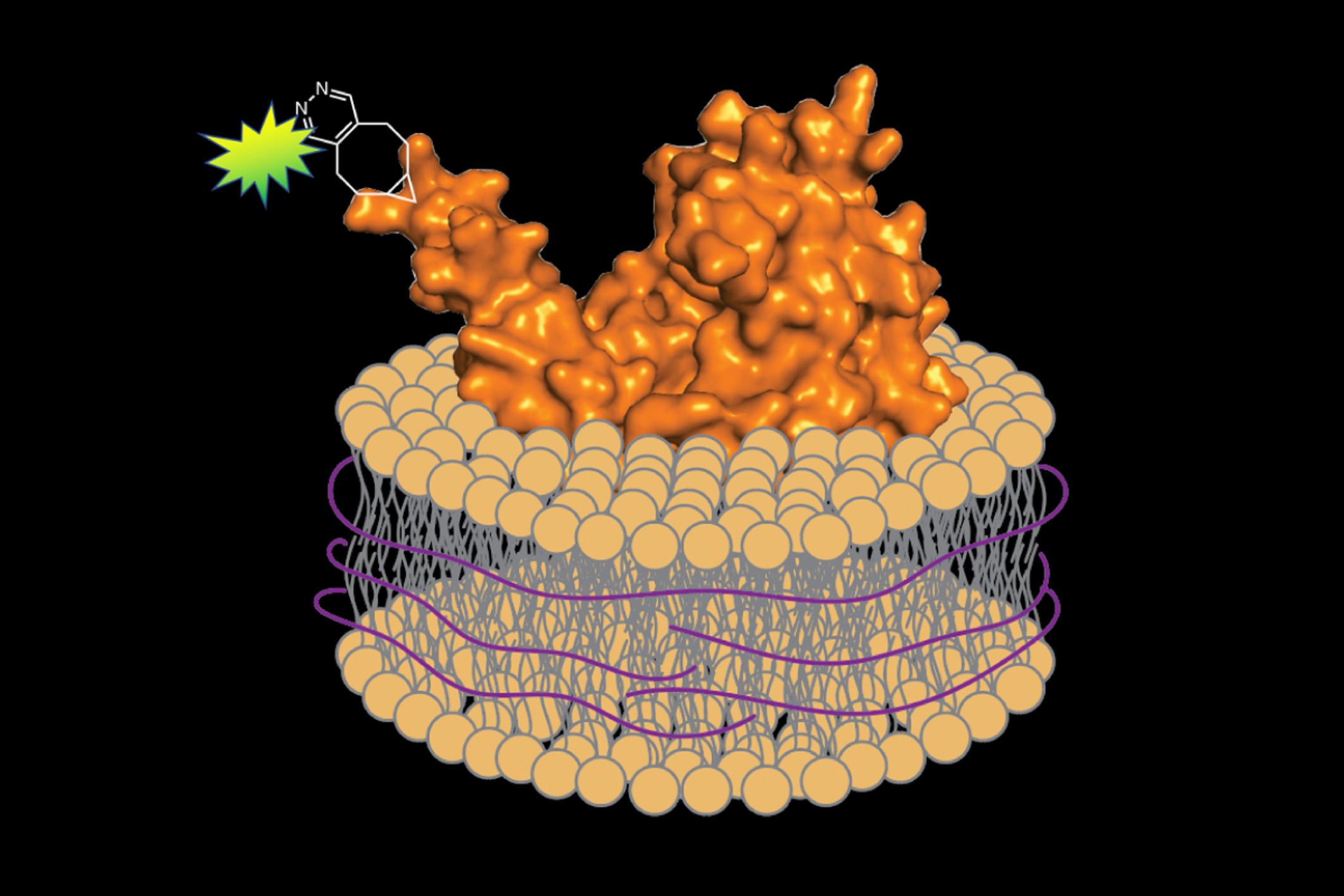
When it came time to remove the proteins from the membrane, he substituted a different substance for the detergent: a polymer of styrene-maleic acid (SMA). Unlike detergent, SMA wraps the extracted protein and a small segment of the associated membrane in a protective shell, preserving its native environment. Imperiali explains, “It’s like a scarf protecting your neck from the cold.”
Swiecicki could then monitor the glowing proteins under a microscope to verify his technique was selective enough to isolate individual membrane proteins. The entire process, he says, takes just a few days, and is generally much faster and more reliable than detergent-based extraction methods, which can take months and require the expertise of highly-trained biochemists to optimize.
“I wouldn’t say it’s a magic bullet that’s going to work for every single protein,” he says. “But it’s a highly efficient tool that could make it easier to study many different kinds of membrane proteins.” Eventually, he says, it may even help facilitate high throughput drug screens.
“As someone who works on membrane protein complexes, I can attest to the great need for better methods to study them,” says Suzanne Walker, a professor of microbiology at Harvard Medical School who was not involved in the study. She hopes to extend the approach outlined in the paper to the protein complexes she investigates in her own lab. “I appreciated the extensive detail included in the text about how to apply the strategy successfully,” she adds.
The next steps will be testing the technique on mammalian proteins, and isolating multiple proteins at once in the SMA shell to observe their interactions. And, of course, every new technique deserves a name. “We’re still working on a catchy acronym,” Imperiali says. “Any ideas?”
This research was funded by the Jane Coffin Childs Memorial Fund for Medical Research, Philippe Foundation, and National Institutes of Health.